¶ What is DataSHIELD?
DataSHIELD is a scientifically mature, open-source federated analysis framework. It enables the remote and privacy-preserving analysis of sensitive research data held across multiple institutions without any individual-level data ever leaving the organisation that holds it. It has been used with real-world research data for over ten years.
Federated analysis: a method of remotely analysing data held across multiple locations without moving or pooling it.
¶ The core idea
Researchers who want to study rare diseases, complex exposures, or large populations often need to combine data from multiple sources (e.g. birth cohort data, clinical records). Traditionally this meant either pooling data in one place — creating practical, legal, and ethical challenges — or running separate analyses at each site and combining results manually.
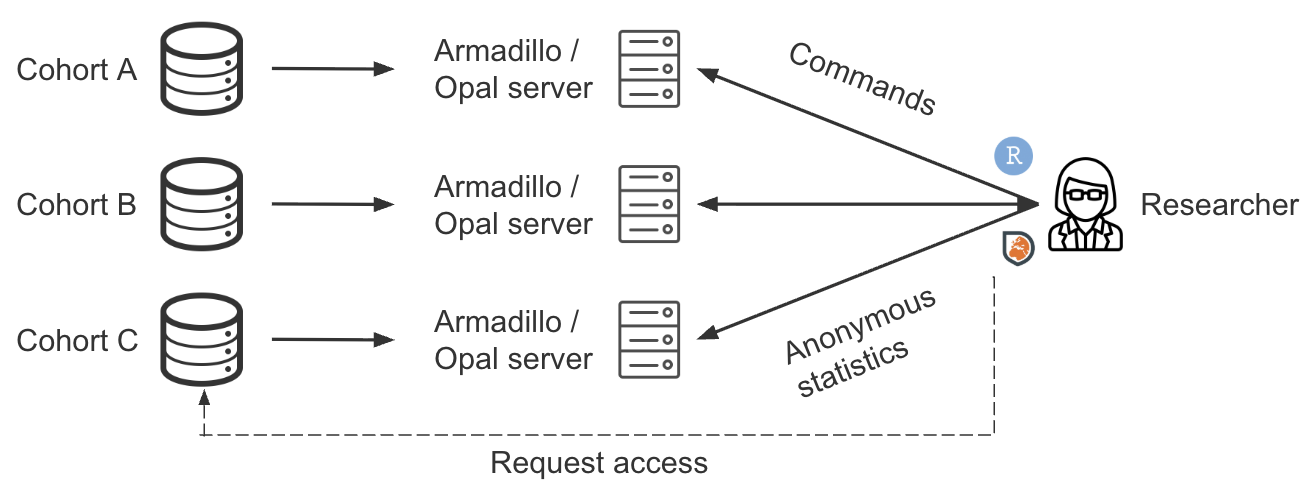
DataSHIELD takes a third path: analysis is brought to the data, not the data to the analyst. A central analyst sends analysis commands to each remote server. Each server runs the computation locally and returns only non-disclosive summary statistics. These summaries are then automatically combined into a single pooled result.
Example: A researcher wants to study the association between air pollution exposure and asthma in children across five European birth cohorts. The data cannot be pooled due to differing national consent frameworks. Using DataSHIELD, the researcher connects to all five cohorts simultaneously and fits a single regression model across 50,000 participants — without any individual-level data leaving its country of origin.
¶ How it works — step by step
-
Data stays local. Each participating institution loads their harmonised dataset onto a server (Armadillo or Opal) running the DataSHIELD server package . No data is transferred to the analyst.
-
The analyst connects remotely. Using the R client package
DSI, an authorised analyst logs in to each server and issues analysis commands from a single R session. Analysts do not connect directly to data sites — access is only available through the DataSHIELD client. -
Commands run in parallel. Each server executes the same analysis on its own slice of data simultaneously. No server can see another server's data.
-
Outputs are checked automatically. Server-side functions are designed to return only non-disclosive summary statistics. Built-in automated output checking runs in real time, based on statistical disclosure control principles. Threshold values are set by the data custodian, not the analyst.
-
Only summaries return. Servers send back aggregate statistics — means, counts, model coefficients — never individual rows. Individual patient data is never viewed by the analyst and is not shared or moved from the data site.
-
Results are pooled. The client-side functions combine the per-server summaries into a combined estimate, either through 'virtual pooling' or two-stage meta-analysis.
¶ How does DataSHIELD differ from other federated platforms?
Several platforms support federated analysis of sensitive data, for example Flower and Vantage6. These follow a batch submission model: the analyst writes a complete algorithm or pipeline, packages it as a Docker container, and submits it to run at each data node. This is well suited to machine learning workflows on quality-controlled data. Disclosure control in this model is not built into individual functions — instead, the completed pipeline is reviewed and approved before deployment.
Other tools like OHDSI and OpenSAFELY work similarly on a job submission basis. Often researchers will have access to a synthetic or a single real world dataset to draft their analysis scripts. Scripts are submitted to a trusted job schedular to run at all data locations, scripts are then checked and run by each data controller the job is submitted to (equivalent to 2-stage analysis, Study Level Meta-analysis). Under these federated tools, outputs are checked by the data controller at each location before being released to the analyst.
DataSHIELD works differently. The analyst works interactively, issuing commands one at a time from an R session and inspecting results as they come back, which fits naturally with the iterative way analysis is typically conducted in epidemiology and biomedical research. Disclosure control is built into every individual function rather than applied at the pipeline level, with the nfilter system automatically enforcing limits on outputs across all analyses.
This makes DataSHIELD highly scalable for multi-cohort research: once a data node is set up and access is granted, any authorised analyst can run analyses without requiring pipeline review or custom deployment. The same R code runs identically whether connecting to two cohorts or twenty, and the disclosure protections apply uniformly across all nodes without additional configuration.
¶ What types of analysis can you do with DataSHIELD?
DataSHIELD's functionality is split across a core package and a growing ecosystem of community extension packages.
dsBase provides the foundation: descriptive statistics, GLMs, linear and generalised linear mixed-effects models, GAMLSS, multiple imputation, data reshaping, and utility functions. dsTidyverse adds tidyverse-style data manipulation, and dsResource enables connections to different resource types including databases and remote files.
Beyond the core, community packages extend DataSHIELD into more specialised domains: survival analysis (dsSurvival), causal mediation (dsMediation), machine learning including PCA, k-means, LASSO and multi-task learning (dsML, dsMTL), genomics and multi-omics (dsOmics), microbiome analysis (dsMicrobiome), exposome research (dsExposome), and generation of synthetic datasets server-side (dsSynthetic).
For a full list see the DataSHIELD community packages page.
¶ Key features
¶ Privacy by design
DataSHIELD follows privacy-by-design principles built into every layer: the computing infrastructure, the analytic functions, and the automated output checking. It is designed to operate with best-practice data governance for the use of sensitive health data.
¶ Federated analysis
Analysis can be aggregated from individual-level data at each location, or a global model can be fitted simultaneously across all sites, without sharing or moving any individual patient data.
¶ Automated output checking
All DataSHIELD analytic functions are designed to output only summary statistics that are low-dimensional and not directly disclosive. Output checking runs automatically in real time; only data custodians can set the threshold values for these checks.
¶ Software validation
The base server package dsBase runs over 12,000 automated tests on every software release. These tests run against synthetic datasets included in the package, checking for correct operation, correct outputs, and that disclosure control is not breached. Armadillo and Opal (the DataSHIELD servers) run their own suites of tests and conduct penetration tests to ensure data security.
¶ Open source
DataSHIELD is a collection of open-source infrastructure and R packages, developed and maintained by the DataSHIELD community.
¶ Key components
| Component | Role |
|---|---|
dsBaseClient |
R package used by analysts. Contains client-side functions that send computation requests to servers. |
dsBase |
Server-side R package. Executes computations on local data and enforces disclosure controls. |
DSI |
DataSHIELD Interface — the R package providing the login and connection layer between client and servers. |
Opal |
OBiBa's data repository and DataSHIELD server backend. |
Armadillo |
MOLGENIS's light-weight alternative DataSHIELD server backend. |
¶ Disclosure control
DataSHIELD's privacy guarantee rests on server-side disclosure filters called nfilters. These are configurable by data custodians and block any result that could identify an individual.
| Filter | What it controls | Default |
|---|---|---|
nfilter.tab |
Minimum cell count in a contingency table | 3 |
nfilter.subset |
Minimum number of individuals in any subset | 3 |
nfilter.glm |
Max model parameters as a proportion of sample size | 0.33 |
nfilter.kNN |
Minimum k for nearest-neighbour operations | 3 |
nfilter.noise |
Minimum noise variance added to returned vectors | 0.25 |
nfilter.levels.density |
Max proportion of unique factor levels regarded as non-disclosive | 0.33 |
nfilter.levels.max |
Max number of unique factor levels regarded as non-disclosive | 40 |
See Disclosure Control for the full reference.
¶ Who governs the DataSHIELD Community?
The DataSHIELD research project is administered by the University of Liverpool. The DataSHIELD Community is a voluntary association whose members include data subjects, data custodians, analysts, statisticians, and developers. A Steering Committee, established in 2023, oversees governance of the open-science, open source DataSHILED community alongside theme leads covering infrastructure, operational management, statistical development, governance, communications, and education.
The project has been under active development since 2009 when it was established by the DataSHIELD research project. Software development is collaborative endeavor, with development and maintenance of core software in the ecosystem provided by Arjuna Technologies, Obiba, the Genomics Coordination Centre (University Medical Centre Groningen), the DataSHIELD research project (University of Liverpool), and other independent contributors.
Full governance details — including the Constitution, voting membership criteria, and the Code of Conduct — are available in the About section.
¶ Further reading
- Beginners tutorial — step-by-step walkthrough of running your first DataSHIELD analysis in R
- Guide for new deployers — information for data custodians considering deploying DataSHIELD
- Disclosure control — full reference for all nfilter settings and privacy control levels
- Publications — searchable index of DataSHIELD research publications
- Community & governance — constitution, voting membership, code of conduct, and how to get involved
- DataSHIELD on GitHub — source code for all packages
- DataSHIELD YouTube channel — introductory videos, webinars, and conference talks